ESGF Compute Function Service Demo
Overview
Prior to this demo, a globus-compute function was registered at the Argonne Leadership Computing Facility (ALCF), which has direct file-access to several petabytes of ESGF data.
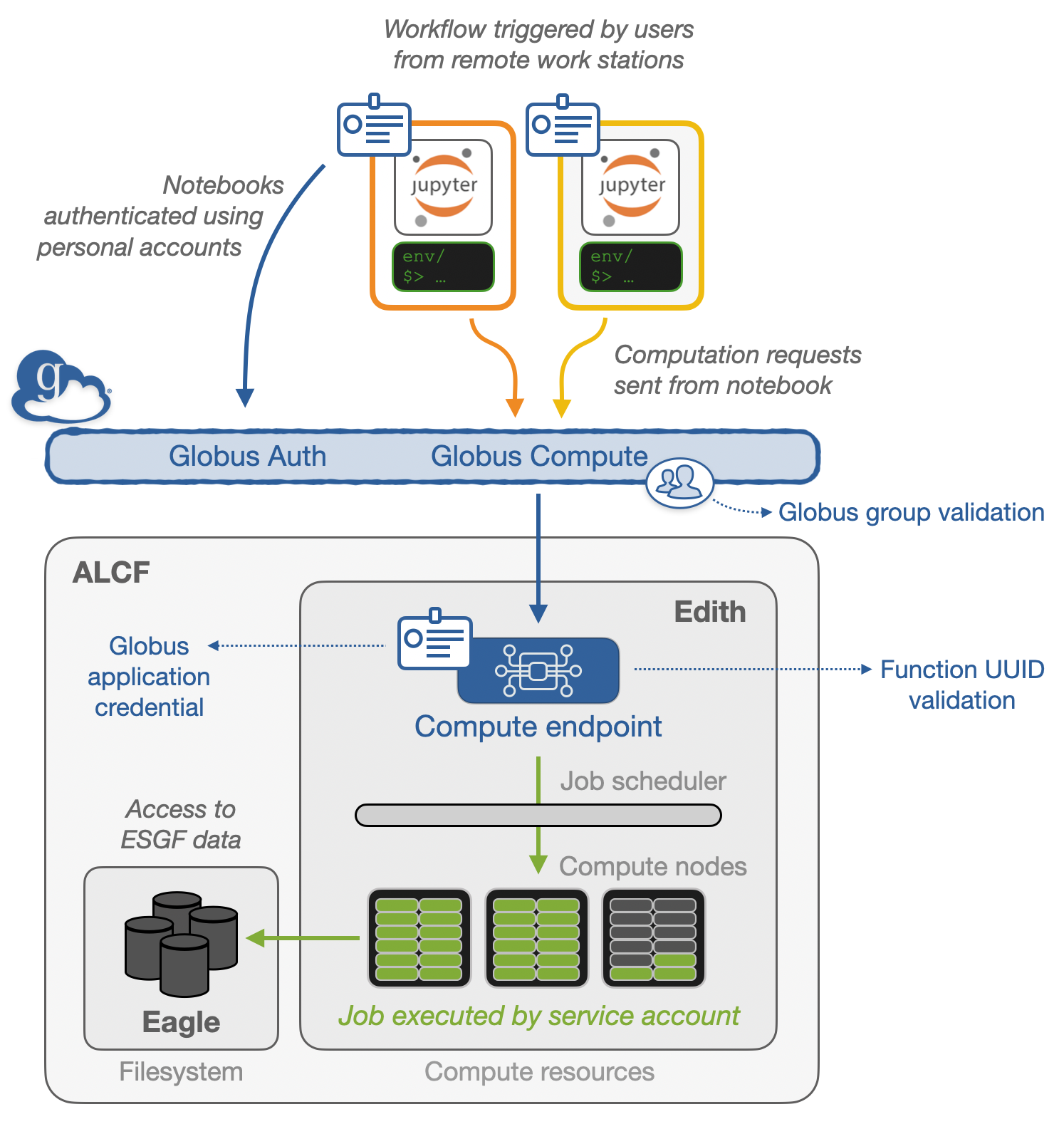
Prerequisites
Concepts |
Importance |
Notes |
|---|---|---|
Necessary |
||
Necessary |
Understanding of globus compute workflows |
|
Helpful |
Familiarity with metadata structure |
|
Helpful |
Familiarity with plotting with Xarray and hvPlot |
Time to learn: 15 minutes
Imports
We need to import a few libraries to visualize the output of our data! The rest of the libraries are installed and run on where we defined the function (ALCF).
# Import packages to help with data visualization
import holoviews as hv
import hvplot
import hvplot.xarray
hv.extension('bokeh')
# Import Globus tools
from globus_compute_sdk import Executor
Remotely Execute + Access ENSO Data from CMIP6
Within this demo, we are looking at a pre-defined, vetted function with the function ID of 49cd1ee0-2c4c-45f1-ab78-c4557fa25aa3. For more on globus-compute function registration, please see the globus compute with ENSO example.
For this function, it takes the source_id as an input, one of the facets from CMIP6.
The result is an xarray.Dataset! The aggregation + computation were done on the high-performance computing cluster, only returning the much smaller dataset.
Create Globus compute executor
Make sure you only define the executor once (unless the executor becomes disconnected and needs a restart).
# Define the UUID of the pre-canned ESGF "run_plot_enso" function
run_plot_enso_function_uuid = "49cd1ee0-2c4c-45f1-ab78-c4557fa25aa3"
# Create Globus Compute executor to run computations at ALCF
endpoint_uuid = "cfaf0e98-2ef3-4c5a-9f11-38e306ddbc2e"
gce = Executor(endpoint_id=endpoint_uuid)
gce.amqp_port = 443
gce
Pass in the source_id of Interest
Source IDs available:
ACCESS-ESM1-5
EC-Earth3-CC
MPI-ESM1-2-LR
CanESM5
MIROC6
EC-Earth3
CESM2
EC-Earth3-Veg
NorCPM1
# Select the target source ID
source_id = "MIROC6"
# Trigger remote computation
future = gce.submit_to_registered_function(run_plot_enso_function_uuid, {source_id})
# Wait for result and generate plot from data returned by the Globus Compute executor
ds = future.result()
ds
encountered unknown data fields while reading a result message: {'details'}
<xarray.Dataset> Size: 77kB
Dimensions: (time: 1980)
Coordinates:
* time (time) datetime64[ns] 16kB 1850-01-16T12:00:00 ... 201...
type (time) |S3 6kB b'sea' b'sea' b'sea' ... b'sea' b'sea'
month (time) int64 16kB 1 2 3 4 5 6 7 8 ... 5 6 7 8 9 10 11 12
Data variables:
tos (time) float32 8kB nan nan -0.3451 ... -1.865 nan nan
tos_gt_04 (time) float32 8kB 0.4 0.4 0.4 0.4 ... 0.4 0.4 0.4 0.4
tos_lt_04 (time) float32 8kB -0.4 -0.4 -0.4 ... -1.865 -0.4 -0.4
el_nino_threshold (time) float32 8kB 0.4 0.4 0.4 0.4 ... 0.4 0.4 0.4 0.4
la_nina_threshold (time) float32 8kB -0.4 -0.4 -0.4 -0.4 ... -0.4 -0.4 -0.4
Attributes: (12/45)
Conventions: CF-1.7 CMIP-6.2
activity_id: CMIP
branch_method: standard
branch_time_in_child: 0.0
branch_time_in_parent: 0.0
creation_date: 2018-11-30T16:23:03Z
... ...
variable_id: tos
variant_label: r1i1p1f1
license: CMIP6 model data produced by MIROC is licensed un...
cmor_version: 3.3.2
tracking_id: hdl:21.14100/31c7618d-6a92-400e-8874-c1fbe41abd44
model: MIROC6Visualize the Dataset Locally
Now that we have the dataset, we can use holoviz tools to create an interactive plot!
def plot_enso(ds):
el_nino = ds.hvplot.area(x="time", y2='tos_gt_04', y='el_nino_threshold', color='red', hover=False)
el_nino_label = hv.Text(ds.isel(time=40).time.values, 2, 'El Niño').opts(text_color='red',)
# Create the La Niña area graphs
la_nina = ds.hvplot.area(x="time", y2='tos_lt_04', y='la_nina_threshold', color='blue', hover=False)
la_nina_label = hv.Text(ds.isel(time=-40).time.values, -2, 'La Niña').opts(text_color='blue')
# Plot a timeseries of the ENSO 3.4 index
enso = ds.tos.hvplot(x='time', line_width=0.5, color='k', xlabel='Year', ylabel='ENSO 3.4 Index')
# Combine all the plots into a single plot
return (el_nino_label * la_nina_label * el_nino * la_nina * enso).opts(title=f'{ds.attrs["model"]} {ds.attrs["source_id"]} \n Ensemble Member: {ds.attrs["variant_label"]}')
plot_enso(ds)
What was in the 49cd1ee0-2c4c-45f1-ab78-c4557fa25aa3 function??
Below is the exact code that was registered at ALCF. If you are interested in writing your own compute functions, please review the Globus Compute with ENSO Notebook
Show code cell source
def run_plot_enso(source_id):
from intake_esgf.exceptions import NoSearchResults
import numpy as np
import matplotlib.pyplot as plt
from intake_esgf import ESGFCatalog
import xarray as xr
import cf_xarray
import warnings
warnings.filterwarnings("ignore")
# List of available source ids
valid_source_id = [
'ACCESS-ESM1-5', 'EC-Earth3-CC', 'MPI-ESM1-2-LR', 'CanESM5',
'MIROC6', 'EC-Earth3', 'CESM2', 'EC-Earth3-Veg', 'NorCPM1'
]
# Validate user input
if not isinstance(source_id, str):
raise ValueError("Source ID should be a string.")
if not source_id in valid_source_id:
raise NoSearchResults("Please use one of the following: "+", ".join(valid_source_id))
def search_esgf(source_id):
# Search and load the ocean surface temperature (tos)
cat = ESGFCatalog(esgf1_indices="anl-dev")
cat.search(
activity_id="CMIP",
experiment_id="historical",
variable_id=["tos"],
source_id=source_id,
member_id='r1i1p1f1',
grid_label="gn",
table_id="Omon",
)
try:
tos_ds = cat.to_dataset_dict()["tos"]
except ValueError:
print(f"Issue with {institution_id} dataset")
return tos_ds
def calculate_enso(ds):
# Subset the El Nino 3.4 index region
dso = ds.where(
(ds.cf["latitude"] < 5) & (ds.cf["latitude"] > -5) & (ds.cf["longitude"] > 190) & (ds.cf["longitude"] < 240), drop=True
)
# Calculate the monthly means
gb = dso.tos.groupby('time.month')
# Subtract the monthly averages, returning the anomalies
tos_nino34_anom = gb - gb.mean(dim='time')
# Determine the non-time dimensions and average using these
non_time_dims = set(tos_nino34_anom.dims)
non_time_dims.remove(ds.tos.cf["T"].name)
weighted_average = tos_nino34_anom.weighted(ds["areacello"].fillna(0)).mean(dim=list(non_time_dims))
# Calculate the rolling average
rolling_average = weighted_average.rolling(time=5, center=True).mean()
std_dev = weighted_average.std()
return rolling_average / std_dev
def add_enso_thresholds(da, threshold=0.4):
# Conver the xr.DataArray into an xr.Dataset
ds = da.to_dataset()
# Cleanup the time and use the thresholds
try:
ds["time"]= ds.indexes["time"].to_datetimeindex()
except:
pass
ds["tos_gt_04"] = ("time", ds.tos.where(ds.tos >= threshold, threshold).data)
ds["tos_lt_04"] = ("time", ds.tos.where(ds.tos <= -threshold, -threshold).data)
# Add fields for the thresholds
ds["el_nino_threshold"] = ("time", np.zeros_like(ds.tos) + threshold)
ds["la_nina_threshold"] = ("time", np.zeros_like(ds.tos) - threshold)
return ds
ds = search_esgf(source_id)
enso_index = add_enso_thresholds(calculate_enso(ds).compute())
enso_index.attrs = ds.attrs
enso_index.attrs["model"] = source_id
return enso_index
Summary
Within this demonstration, we remotely triggered a globus-compute function which read, aggregated, and computed ENSO on datasets located on an HPC system. The computations were all done on the server side, with visualization being the only task done locally.
What’s next?
Some existing questions still exist! Mainly:
Where do define these functions? How do we ensure these are safe to run?
How do we request “service” accounts on other HPC/cloud facilities?
What other functions, outside of the typical WPS services, can we define?
What other use-cases could this support (ex. kerchunk or zarr creation)?